Section 1: Introduction¶
This notebook illustrates how to compute and plot various Nino indices using the HadISST dataset hosted on NCAR’s Geoscience Data Exchange (GDEX)
Python package imports and useful function definitions
# Import
import intake
import numpy as np
import xarray as xr
import nc_time_axis
import os
import matplotlib.pyplot as pltimport dask
from dask_jobqueue import PBSCluster
from dask.distributed import Client
from dask.distributed import performance_report# Set up your sratch folder path
username = os.environ["USER"]
glade_scratch = "/glade/derecho/scratch/" + username
print(glade_scratch)/glade/derecho/scratch/harshah
#### Useful function ###
def area_mean(da):
"""
Area-weighted mean over lat/lon using cos(lat) weighting.
Assumes da has dims including lat, lon.
"""
weights = np.cos(np.deg2rad(da["latitude"]))
weights.name = "weights"
return da.weighted(weights).mean(dim=("latitude", "longitude"), skipna=True)
def lon_to_360(da):
"""Ensure longitude is [0, 360) and sorted."""
lon_name = "lon" if "lon" in da.coords else "longitude"
lon = da[lon_name]
lon360 = (lon % 360)
#The above logic is tricky for negative divisors or dividends and is not guaranteed to work in other programming languages
#For more details, see https://en.wikipedia.org/wiki/Modulo
da = da.assign_coords({lon_name: lon360}).sortby(lon_name)
return da##
def nino_index_series(sst, region):
lat0, lat1 = region["latitude"]
lon0, lon1 = region["longitude"]
sub = sst.sel(latitude=slice(lat0, lat1), longitude=slice(lon0, lon1))
return area_mean(sub)Section 2: Set up a Dask Cluster on Casper¶
Setting up a dask cluster. You will need an NCAR HPC account to do this.
# Create a PBS cluster object
cluster = PBSCluster(
job_name = 'dask-wk25',
cores = 1,
memory = '8GiB',
processes = 1,
local_directory = glade_scratch+'/dask/spill/',
log_directory = glade_scratch + '/dask/logs/',
resource_spec = 'select=1:ncpus=1:mem=8GB',
queue = 'casper',
walltime = '5:00:00',
interface = 'ext'
)# Create the client to load the Dashboard
client = Client(cluster)n_workers = 5
cluster.scale(n_workers)
client.wait_for_workers(n_workers = n_workers)
clusterSection 3: Data Loading and transforms¶
Load HadISST data from GDEX using xarray
For more details regarding the dataset. See, https://
gdex .ucar .edu /datasets /d27703 /#
import gzip
import tempfile
with gzip.open("/gdex/data/d277003/HadISST_sst.nc.gz", "rb") as f:
raw = f.read()
with tempfile.NamedTemporaryFile(suffix=".nc") as tmp:
tmp.write(raw)
tmp.flush()
ds = xr.open_dataset(tmp.name)sst_raw = ds.sst
sst_rawConvert longitude from [-180,180) to [0,360)
sst = lon_to_360(sst_raw)
sstSection 4: Data Analysis¶
Define the various nino regions and their spatial extent as a dictionary
Compute area weighted spatial mean
Make a quick plot of Nino indices
Compute anomalies w.r.t to a baseline period
Compute 6 month running mean and visualize El Nino events
Optional: Compare with NOAA Nino index calculation
NINO_REGIONS = {
"nino12": {"latitude": (0, -10), "longitude": (270, 280)}, # 90W–80W
"nino3": {"latitude": (5, -5), "longitude": (210, 270)}, # 150W–90W
"nino34": {"latitude": (5, -5), "longitude": (190, 240)}, # 170W–120W
"nino4": {"latitude": (5, -5), "longitude": (160, 210)}, # 160E–150W
}nino_raw = xr.Dataset({name: nino_index_series(sst, reg) for name, reg in NINO_REGIONS.items()})
nino_rawnino_raw.to_dataframe().plot(subplots=True, figsize=(10, 8), lw=1)
plt.gcf().supylabel(r'SST ($^\circ$ C)')
plt.gcf().supxlabel('time')
plt.tight_layout()
plt.gcf().suptitle(' Nino indices from monthly HadISST data',fontsize=12,fontweight='bold',y=0.999)
plt.show()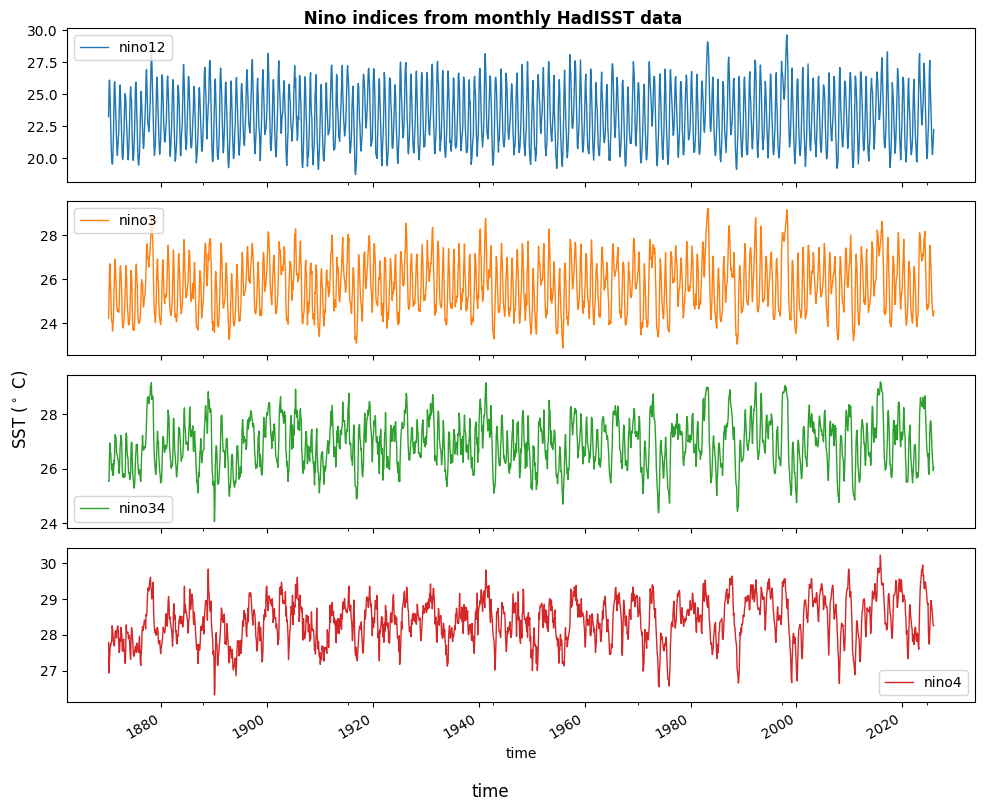
# %%time
# sst = xr.open_dataset('/gdex/data/d277003/HadISST_sst.nc.gz',engine='netcdf4')Compute Nino anomalies¶
Acknowledgements¶
Trenberth, Kevin & National Center for Atmospheric Research Staff (Eds). Last modified 2025-12-11 "The Climate Data Guide: Nino SST Indices (Nino 1+2, 3, 3.4, 4; ONI and TNI).” Retrieved from https://
climatedataguide .ucar .edu /climate -data /nino -sst -indices -nino -12 -3 -34 -4 -oni -and -tni on 2026-02-25.
cluster.close()