This notebook is adapted from the NCAR gallery in the Pangeo collection
Section 1: Introduction¶
Input Data Access¶
This notebook illustrates how to compute surface ocean heat content using potential temperature data from CESM2 Large Ensemble Dataset (https://
www .cesm .ucar .edu /community -projects /lens2) hosted on NCAR’s GDEX. This data is open access and is accessed via OSDF
# Imports
import intake
import numpy as np
import pandas as pd
import xarray as xr
import seaborn as sns
import re
import os
import matplotlib.pyplot as pltimport fsspec.implementations.http as fshttp
from pelicanfs.core import PelicanFileSystem, PelicanMap, OSDFFileSystem
import cf_units as cfimport dask
from dask_jobqueue import PBSCluster
from dask.distributed import Client
from dask.distributed import performance_reportinit_year0 = '1991'
init_year1 = '2020'
final_year0 = '2071'
final_year1 = '2100'def to_daily(ds):
year = ds.time.dt.year
day = ds.time.dt.dayofyear
# assign new coords
ds = ds.assign_coords(year=("time", year.data), day=("time", day.data))
# reshape the array to (..., "day", "year")
return ds.set_index(time=("year", "day")).unstack("time")# Set up your sratch folder path
username = os.environ["USER"]
glade_scratch = "/glade/derecho/scratch/" + username
print(glade_scratch)
#
catalog_url = 'https://osdata.gdex.ucar.edu/d010092/catalogs/d010092-osdf.json'
# catalog_url = 'https://data.gdex.ucar.edu/OS/d010092/catalogs/d010092-https.json' #NCAR's Object store/glade/derecho/scratch/harshah
Section 2: Select Dask Cluster¶
Setting up a dask cluster. You will need to choose the type of cluster based on your use case.
The default will be LocalCluster as that can run on any system.
If running on NCAR’s HPC i.e, Casper with a PBS Scheduler, set USE_PBS_SCHEDULER to True. Otherwise, set to False.
# Create a PBS cluster object
cluster = PBSCluster(
job_name = 'dask-wk24-hpc',
cores = 1,
memory = '8GiB',
processes = 1,
local_directory = glade_scratch+'/dask/spill',
log_directory = glade_scratch + '/dask/logs/',
resource_spec = 'select=1:ncpus=1:mem=8GB',
queue = 'casper',
walltime = '5:00:00',
#interface = 'ib0'
interface = 'ext'
)/glade/u/home/harshah/.conda/envs/osdf/lib/python3.11/site-packages/distributed/node.py:188: UserWarning: Port 8787 is already in use.
Perhaps you already have a cluster running?
Hosting the HTTP server on port 43825 instead
warnings.warn(
# Create the client to load the Dashboard
client = Client(cluster)n_workers =5
cluster.scale(n_workers)
client.wait_for_workers(n_workers = n_workers)
clusterSection 3: Data Loading¶
Load CESM2 LENS zarr data from GDEX using an intake-ESM catalog
For more details regarding the dataset. See, https://
gdex .ucar .edu /datasets /d010092 /#
cesm_cat = intake.open_esm_datastore(catalog_url)
cesm_cat# cesm_cat.df['variable'].valuescesm_temp = cesm_cat.search(variable ='TEMP', frequency ='monthly',experiment='historical')
cesm_tempcesm_temp.df['path'].values<ArrowExtensionArray>
['osdf:///ncar-gdex/d010092/ocn/monthly/cesm2LE-historical-cmip6-TEMP.zarr']
Length: 1, dtype: large_string[pyarrow]dsets_cesm = cesm_temp.to_dataset_dict(xarray_open_kwargs={'engine':'zarr','backend_kwargs':{'consolidated': True,'zarr_format': 2}})
--> The keys in the returned dictionary of datasets are constructed as follows:
'component.experiment.frequency.forcing_variant'
cesm_temp.keys()['ocn.historical.monthly.cmip6']historical = dsets_cesm['ocn.historical.monthly.cmip6']
# future_smbb = dsets_cesm['ocn.ssp370.monthly.smbb']
# future_cmip6 = dsets_cesm['ocn.ssp370.monthly.cmip6']# %%time
# merge_ds_cmip6 = xr.concat([historical, future_cmip6], dim='time')
# merge_ds_cmip6 = merge_ds_cmip6.dropna(dim='member_id')historicalChange units¶
orig_units = cf.Unit(historical.z_t.attrs['units'])
orig_unitsUnit('centimeters')def change_units(ds, variable_str, variable_bounds_str, target_unit_str):
orig_units = cf.Unit(ds[variable_str].attrs['units'])
target_units = cf.Unit(target_unit_str)
variable_in_new_units = xr.apply_ufunc(orig_units.convert, ds[variable_bounds_str], target_units, dask='parallelized', output_dtypes=[ds[variable_bounds_str].dtype])
return variable_in_new_unitshistorical['z_t']depth_levels_in_m = change_units(historical, 'z_t', 'z_t', 'm')
hist_temp_in_degK = change_units(historical, 'TEMP', 'TEMP', 'degK')
# fut_cmip6_temp_in_degK = change_units(future_cmip6, 'TEMP', 'TEMP', 'degK')
# fut_smbb_temp_in_degK = change_units(future_smbb, 'TEMP', 'TEMP', 'degK')
#
hist_temp_in_degK = hist_temp_in_degK.assign_coords(z_t=("z_t", depth_levels_in_m['z_t'].data))
hist_temp_in_degK["z_t"].attrs["units"] = "m"
hist_temp_in_degKdepth_levels_in_m.isel(z_t=slice(0, -1))#Compute depth level deltas using z_t levels
depth_level_deltas = depth_levels_in_m.isel(z_t=slice(1, None)).values - depth_levels_in_m.isel(z_t=slice(0, -1)).values
# Optionally, if you want to keep it as an xarray DataArray, re-wrap the result
depth_level_deltas = xr.DataArray(depth_level_deltas, dims=["z_t"], coords={"z_t": depth_levels_in_m.z_t.isel(z_t=slice(0, -1))})
depth_level_deltas Section 4: Data Analysis¶
Compute Ocean Heat content for ocean surface¶
Ocean surface is considered to be the top 100m
The formula for this is:
Where H is ocean heat content, the value we are trying to calculate,
is the density of sea water, ,
is the specific heat of sea water, ,
is the depth limit of the calculation in meters,
and is the temperature at each depth in degrees Kelvin.
def calc_ocean_heat(delta_level, temperature):
rho = 1026 #kg/m^3
c_p = 3990 #J/(kg K)
weighted_temperature = delta_level * temperature
heat = weighted_temperature.sum(dim="z_t")*rho*c_p
return heat# Remember that the coordinate z_t still has values in cm
hist_temp_ocean_surface = hist_temp_in_degK.where(hist_temp_in_degK['z_t'] < 1e4,drop=True)
hist_temp_ocean_surfacedepth_level_deltas_surface = depth_level_deltas.where(depth_level_deltas['z_t'] <1e4, drop= True)
depth_level_deltas_surfacehist_ocean_heat = calc_ocean_heat(depth_level_deltas_surface,hist_temp_ocean_surface)
hist_ocean_heatPlot Ocean Heat¶
%%time
# Jan, 1850 average over all memebers
# hist_ocean_avgheat = hist_ocean_heat.mean('member_id')
hist_ocean_avgheat = hist_ocean_heat.isel({'time':[0,-12]}).mean('member_id')
hist_ocean_avgheatCPU times: user 39.6 ms, sys: 3.67 ms, total: 43.3 ms
Wall time: 42.9 ms
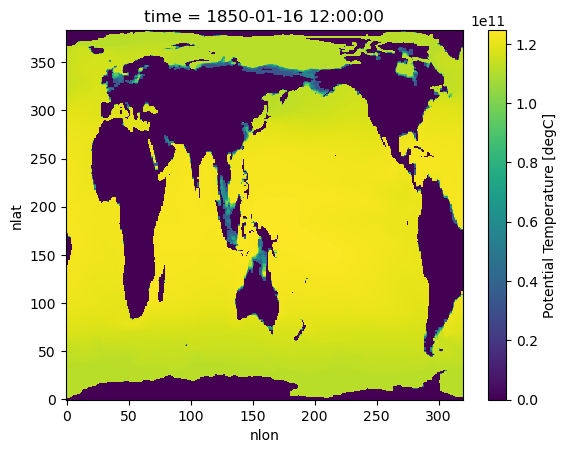
%%time
hist_ocean_avgheat.isel(time=0).plot()/glade/u/home/harshah/.conda/envs/osdf/lib/python3.11/site-packages/distributed/client.py:3375: UserWarning: Sending large graph of size 14.27 MiB.
This may cause some slowdown.
Consider loading the data with Dask directly
or using futures or delayed objects to embed the data into the graph without repetition.
See also https://docs.dask.org/en/stable/best-practices.html#load-data-with-dask for more information.
warnings.warn(
CPU times: user 11.5 s, sys: 687 ms, total: 12.2 s
Wall time: 1min 30s
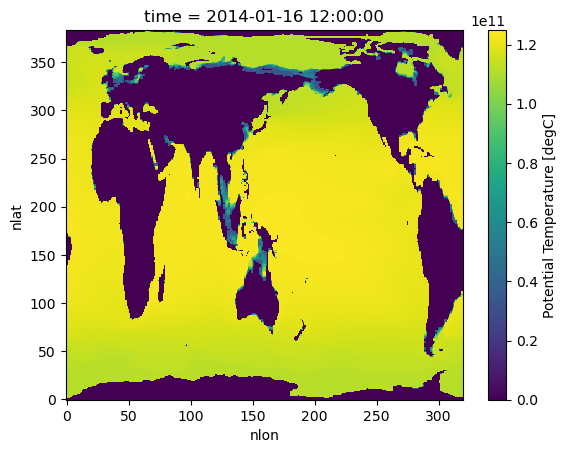
%%time
#Plot ocean heat for Jan 2014
hist_ocean_avgheat.isel(time=1).plot()CPU times: user 11.5 s, sys: 455 ms, total: 12 s
Wall time: 1min 3s
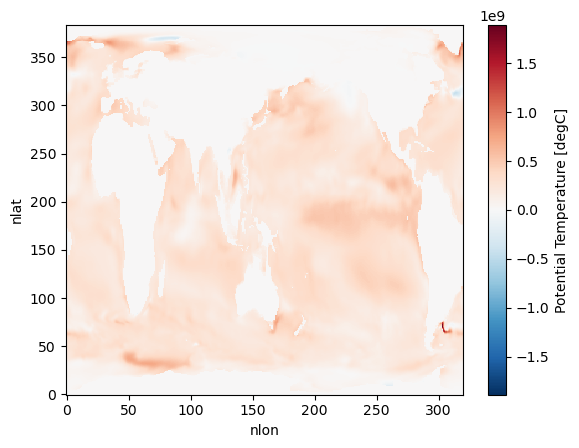
Has the surface ocean heat content increased with time for January ? (Due to Global Warming!)¶
hist_ocean_avgheat_ano = hist_ocean_avgheat.isel(time=1) - hist_ocean_avgheat.isel(time=0)%%time
hist_ocean_avgheat_ano.plot()/glade/u/home/harshah/.conda/envs/osdf/lib/python3.11/site-packages/distributed/client.py:3375: UserWarning: Sending large graph of size 14.27 MiB.
This may cause some slowdown.
Consider loading the data with Dask directly
or using futures or delayed objects to embed the data into the graph without repetition.
See also https://docs.dask.org/en/stable/best-practices.html#load-data-with-dask for more information.
warnings.warn(
CPU times: user 10.9 s, sys: 344 ms, total: 11.2 s
Wall time: 1min 1s
cluster.close()