Access CMIP6 zarr data from AWS using the osdf protocol and compute GMSTA on a Jetstream2 Exosphere instance¶
This workflow is inspired by https://
gallery .pangeo .io /repos /pangeo -gallery /cmip6 /global _mean _surface _temp .html
Table of Contents¶
Section 1: Introduction¶
Load python packkages
Load catalog url
from matplotlib import pyplot as plt
import xarray as xr
import numpy as np
from dask.diagnostics import progress
from tqdm.autonotebook import tqdm
import intake
import fsspec
import seaborn as sns
import aiohttp
import dask
from dask.distributed import LocalCluster
import pelicanfs
import os
from pelicanfs.core import OSDFFileSystem,PelicanMap /tmp/ipykernel_3608/4282021079.py:5: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from tqdm.autonotebook import tqdm
#
gdex_url = 'https://data.gdex.ucar.edu/'
cat_url = gdex_url + 'd850001/catalogs/osdf/cmip6-aws/cmip6-osdf-zarr.json'Section 2: Select Dask Cluster¶
Setting up a dask cluster.
The default will be LocalCluster as that can run on any system.
cluster = LocalCluster()
client = cluster.get_client()# Scale the local cluster
n_workers = 5
cluster.scale(n_workers)
clusterLoading...
Section 3: Data Loading¶
Load catalog and select data subset
col = intake.open_esm_datastore(cat_url)
colLoading...
[eid for eid in col.df['experiment_id'].unique() if 'ssp' in eid]['esm-ssp585-ssp126Lu',
'ssp126-ssp370Lu',
'ssp370-ssp126Lu',
'ssp585',
'ssp245',
'ssp370-lowNTCF',
'ssp370SST-ssp126Lu',
'ssp370SST',
'ssp370pdSST',
'ssp370SST-lowCH4',
'ssp370SST-lowNTCF',
'ssp126',
'ssp119',
'ssp370',
'esm-ssp585',
'ssp245-nat',
'ssp245-GHG',
'ssp460',
'ssp434',
'ssp534-over',
'ssp245-aer',
'ssp245-stratO3',
'ssp245-cov-fossil',
'ssp245-cov-modgreen',
'ssp245-cov-strgreen',
'ssp245-covid',
'ssp585-bgc']# there is currently a significant amount of data for these runs
expts = ['historical', 'ssp245', 'ssp370']
query = dict(
experiment_id=expts,
table_id='Amon',
variable_id=['tas'],
member_id = 'r1i1p1f1',
#activity_id = 'CMIP',
)
col_subset = col.search(require_all_on=["source_id"], **query)
col_subsetLoading...
col_subset.df.groupby("source_id")[["experiment_id", "variable_id", "table_id","activity_id"]].nunique()Loading...
col_subset.dfLoading...
# %%time
# dsets_osdf = col_subset.to_dataset_dict()
# print(f"\nDataset dictionary keys:\n {dsets_osdf.keys()}")def drop_all_bounds(ds):
drop_vars = [vname for vname in ds.coords
if (('_bounds') in vname ) or ('_bnds') in vname]
return ds.drop_vars(drop_vars)
def open_dset(df):
assert len(df) == 1
mapper = fsspec.get_mapper(df.zstore.values[0])
path = df.zstore.values[0][7:]+".zmetadata"
ds = xr.open_zarr(mapper, consolidated=True,zarr_format=2) #You can skip the zarr_ormat option if you have zarr version 2
a_data = mapper.fs.get_access_data()
responses = a_data.get_responses(path)
print(responses)
return drop_all_bounds(ds)
def open_delayed(df):
return dask.delayed(open_dset)(df)
from collections import defaultdict
dsets = defaultdict(dict)
for group, df in col_subset.df.groupby(by=['source_id', 'experiment_id']):
dsets[group[0]][group[1]] = open_delayed(df)dsets_ = dask.compute(dict(dsets))[0]#calculate global means
def get_lat_name(ds):
for lat_name in ['lat', 'latitude']:
if lat_name in ds.coords:
return lat_name
raise RuntimeError("Couldn't find a latitude coordinate")
def global_mean(ds):
lat = ds[get_lat_name(ds)]
weight = np.cos(np.deg2rad(lat))
weight /= weight.mean()
other_dims = set(ds.dims) - {'time'}
return (ds * weight).mean(other_dims)expt_da = xr.DataArray(expts, dims='experiment_id', name='experiment_id',
coords={'experiment_id': expts})
dsets_aligned = {}
for k, v in tqdm(dsets_.items()):
expt_dsets = v.values()
if any([d is None for d in expt_dsets]):
print(f"Missing experiment for {k}")
continue
for ds in expt_dsets:
ds.coords['year'] = ds.time.dt.year
# workaround for
# https://github.com/pydata/xarray/issues/2237#issuecomment-620961663
dsets_ann_mean = [v[expt].pipe(global_mean).swap_dims({'time': 'year'})
.drop_vars('time').coarsen(year=12).mean()
for expt in expts]
# align everything with the 4xCO2 experiment
dsets_aligned[k] = xr.concat(dsets_ann_mean, join='outer',dim=expt_da)100%|███████████████████████████████████████████████████████████████████████████████| 27/27 [00:04<00:00, 6.00it/s]
%%time
with progress.ProgressBar():
dsets_aligned_ = dask.compute(dsets_aligned)[0]CPU times: user 12.2 s, sys: 1.85 s, total: 14 s
Wall time: 1min 46s
source_ids = list(dsets_aligned_.keys())
source_da = xr.DataArray(source_ids, dims='source_id', name='source_id',
coords={'source_id': source_ids})
big_ds = xr.concat([ds.reset_coords(drop=True)
for ds in dsets_aligned_.values()],
dim=source_da)
big_ds/tmp/ipykernel_3608/2886188115.py:5: FutureWarning: In a future version of xarray the default value for join will change from join='outer' to join='exact'. This change will result in the following ValueError: cannot be aligned with join='exact' because index/labels/sizes are not equal along these coordinates (dimensions): 'year' ('year',) The recommendation is to set join explicitly for this case.
big_ds = xr.concat([ds.reset_coords(drop=True)
Loading...
Observational time series data for comparison with ensemble spread¶
The observational data we will use is the HadCRUT5 dataset from the UK Met Office
The data has been downloaded to NCAR’s Geoscience Data Exchange (GDEX) from https://
www .metoffice .gov .uk /hadobs /hadcrut5/ We will use an OSDF file system to access this copy from the GDEX
osdf_fs = OSDFFileSystem()
print(osdf_fs)
#
obs_url = '/ncar/gdex/d850001/HadCRUT.5.0.2.0.analysis.summary_series.global.monthly.nc'
#
obs_ds = xr.open_dataset(osdf_fs.open(obs_url,mode='rb'),engine='h5netcdf').tas_mean
obs_ds<pelicanfs.core.OSDFFileSystem object at 0x744ff5790b50>
Loading...
# Compute annual mean temperatures anomalies
obs_gmsta = obs_ds.resample(time='YS').mean(dim='time')
obs_gmstaLoading...
Section 4: Data Analysis¶
Calculate Global Mean Surface Temperature Anomaly (GMSTA)
Grab some observations/ ERA5 reanalysis data
Convert xarray datasets to dataframes
Use Seaborn to plot GMSTA
df_all = big_ds.to_dataframe().reset_index()
df_all.head()Loading...
# Compute anomaly w.r.t 1960-1990 baseline
# Define the baseline period
baseline_df = df_all[(df_all["year"] >= 1960) & (df_all["year"] <= 1990)]
# Compute the baseline mean
baseline_mean = baseline_df["tas"].mean()
# Compute anomalies
df_all["tas_anomaly"] = df_all["tas"] - baseline_mean
df_allLoading...
obs_df = obs_gmsta.to_dataframe(name='tas_anomaly').reset_index()
# obs_df# Convert 'time' to 'year' (keeping only the year)
obs_df['year'] = obs_df['time'].dt.year
# Drop the original 'time' column since we extracted 'year'
obs_df = obs_df[['year', 'tas_anomaly']]
obs_dfLoading...
# Create the main plot
g = sns.relplot(data=df_all, x="year", y="tas_anomaly",
hue='experiment_id', kind="line", errorbar="sd", aspect=2, palette="Set2") # Adjust the color palette)
# Get the current axis from the FacetGrid
ax = g.ax
# Overlay the observational data in red
sns.lineplot(data=obs_df, x="year", y="tas_anomaly",color="red",
linestyle="dashed", linewidth=2,label="Observations", ax=ax)
# Adjust the legend to include observations
ax.legend(title="Experiment ID + Observations")
# Show the plot
plt.show()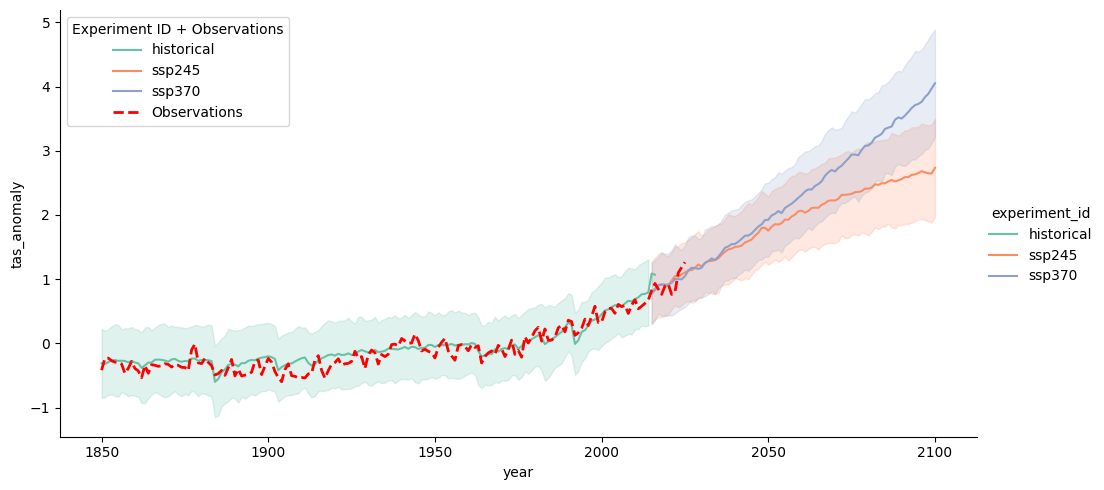
# cluster.close()# sns.relplot(data=df_all,x="year", y="tas_anomaly", hue='experiment_id',kind="line", errorbar="sd", aspect=2)