origin:ncar-posix origin:ncar-object-store platform:casper dataset:na-cordex task:visualization level:intermediate
This notebook is adapted from the NA CORDEX notebook on AWS
http://
ncar -aws -www .s3 -website -us -west -2 .amazonaws .com /plot -zarr -diagnostics .html
Input Data Access¶
This notebook illustrates how to make diagnostic plots using the NA-CORDEX dataset hosted on NCAR’s Geoscience Data Exchange (GDEX)
This data is open access and can be accessed via 2 protocols
HTTPS (if you have access to NCAR’s HPC)
OSDF using an intake-ESM catalog.
# Imports
import intake
import numpy as np
import pandas as pd
import xarray as xr
import seaborn as sns
import re
import matplotlib.pyplot as plt
import dask
from dask_jobqueue import PBSCluster
from dask.distributed import Client
import osinit_year0 = '1991'
init_year1 = '2020'
final_year0 = '2071'
final_year1 = '2100'# catalog_url = 'https://osdf-data.gdex.ucar.edu/ncar-gdex/d316010/catalogs/d316010-osdf-zarr.json'
catalog_url = 'https://osdata.gdex.ucar.edu/d316010/catalogs/d316010-osdf.json' #NCAR's Object store
print(catalog_url)https://osdata.gdex.ucar.edu/d316010/catalogs/d316010-osdf.json
# Set up your scratch folder path
username = os.environ["USER"]
glade_scratch = "/glade/derecho/scratch/" + username
print(glade_scratch)/glade/derecho/scratch/harshah
Create a PBS cluster¶
# Create a PBS cluster object
cluster = PBSCluster(
job_name = 'dask-wk24-hpc',
cores = 1,
memory = '8GiB',
processes = 1,
local_directory = glade_scratch+'/dask/spill',
log_directory = glade_scratch + '/dask/logs/',
resource_spec = 'select=1:ncpus=1:mem=8GB',
queue = 'casper',
walltime = '5:00:00',
#interface = 'ib0'
interface = 'ext'
)/glade/u/home/harshah/.conda/envs/osdf/lib/python3.11/site-packages/distributed/node.py:188: UserWarning: Port 8787 is already in use.
Perhaps you already have a cluster running?
Hosting the HTTP server on port 34989 instead
warnings.warn(
# Create the client to load the Dashboard
client = Client(cluster)
n_workers = 8
cluster.scale(n_workers)
client.wait_for_workers(n_workers = n_workers)
clusterLoading...
Load NA CORDEX data from GDEX using an intake catalog¶
col = intake.open_esm_datastore(catalog_url)
colLoading...
Load data into xarray¶
data_var = 'tmax'
col_subset = col.search(
variable=data_var,
grid="NAM-44i",
scenario="eval",
bias_correction="raw",
)
col_subsetLoading...
col_subset.df['path'].values<ArrowExtensionArray>
['osdf:////ncar-gdex/d316010/day/tmax.eval.day.NAM-44i.raw.zarr']
Length: 1, dtype: large_string[pyarrow]col_subset['tmax.day.eval.NAM-44i.raw'].dfLoading...
# Load catalog entries for subset into a dictionary of xarray datasets, and open the first one.
dsets = col_subset.to_dataset_dict(zarr_kwargs={"consolidated": True,'zarr_format':2})
print(f"\nDataset dictionary keys:\n {dsets.keys()}")
--> The keys in the returned dictionary of datasets are constructed as follows:
'variable.frequency.scenario.grid.bias_correction'
Loading...
Dataset dictionary keys:
dict_keys(['tmax.day.eval.NAM-44i.raw'])
# Load the first dataset and display a summary.
dataset_key = list(dsets.keys())[0]
store_name = dataset_key + ".zarr"
ds = dsets[dataset_key]
dsLoading...
Functions for Plotting¶
Helper Function to Create a Single Map Plot¶
def plotMap(ax, map_slice, date_object=None, member_id=None):
"""Create a map plot on the given axes, with min/max as text"""
ax.imshow(map_slice, origin='lower')
minval = map_slice.min(dim = ['lat', 'lon'])
maxval = map_slice.max(dim = ['lat', 'lon'])
# Format values to have at least 4 digits of precision.
ax.text(0.01, 0.03, "Min: %3g" % minval, transform=ax.transAxes, fontsize=12)
ax.text(0.99, 0.03, "Max: %3g" % maxval, transform=ax.transAxes, fontsize=12, horizontalalignment='right')
ax.set_xticks([])
ax.set_yticks([])
if date_object:
ax.set_title(date_object.values.astype(str)[:10], fontsize=12)
if member_id:
ax.set_ylabel(member_id, fontsize=12)
return axHelper Function for Finding Dates with Available Data¶
def getValidDateIndexes(member_slice):
"""Search for the first and last dates with finite values."""
min_values = member_slice.min(dim = ['lat', 'lon'])
is_finite = np.isfinite(min_values)
finite_indexes = np.where(is_finite)
start_index = finite_indexes[0][0]
end_index = finite_indexes[0][-1]
return start_index, end_indexFunction Producing Maps of First, Middle, and Final Timesteps¶
def plot_first_mid_last(ds, data_var, store_name):
"""Plot the first, middle, and final time steps for several climate runs."""
num_members_to_plot = 4
member_names = ds.coords['member_id'].values[0:num_members_to_plot]
figWidth = 18
figHeight = 12
numPlotColumns = 3
fig, axs = plt.subplots(num_members_to_plot, numPlotColumns, figsize=(figWidth, figHeight), constrained_layout=True)
for index in np.arange(num_members_to_plot):
mem_id = member_names[index]
data_slice = ds[data_var].sel(member_id=mem_id)
start_index, end_index = getValidDateIndexes(data_slice)
midDateIndex = np.floor(len(ds.time) / 2).astype(int)
startDate = ds.time[start_index]
first_step = data_slice.sel(time=startDate)
ax = axs[index, 0]
plotMap(ax, first_step, startDate, mem_id)
midDate = ds.time[midDateIndex]
mid_step = data_slice.sel(time=midDate)
ax = axs[index, 1]
plotMap(ax, mid_step, midDate)
endDate = ds.time[end_index]
last_step = data_slice.sel(time=endDate)
ax = axs[index, 2]
plotMap(ax, last_step, endDate)
plt.suptitle(f'First, Middle, and Last Timesteps for Selected Runs in "{store_name}"', fontsize=20)
return figFunction Producing Statistical Map Plots¶
def plot_stat_maps(ds, data_var, store_name):
"""Plot the mean, min, max, and standard deviation values for several climate runs, aggregated over time."""
num_members_to_plot = 4
member_names = ds.coords['member_id'].values[0:num_members_to_plot]
figWidth = 25
figHeight = 12
numPlotColumns = 4
fig, axs = plt.subplots(num_members_to_plot, numPlotColumns, figsize=(figWidth, figHeight), constrained_layout=True)
for index in np.arange(num_members_to_plot):
mem_id = member_names[index]
data_slice = ds[data_var].sel(member_id=mem_id)
data_agg = data_slice.min(dim='time')
plotMap(axs[index, 0], data_agg, member_id=mem_id)
data_agg = data_slice.max(dim='time')
plotMap(axs[index, 1], data_agg)
data_agg = data_slice.mean(dim='time')
plotMap(axs[index, 2], data_agg)
data_agg = data_slice.std(dim='time')
plotMap(axs[index, 3], data_agg)
axs[0, 0].set_title(f'min({data_var})', fontsize=15)
axs[0, 1].set_title(f'max({data_var})', fontsize=15)
axs[0, 2].set_title(f'mean({data_var})', fontsize=15)
axs[0, 3].set_title(f'std({data_var})', fontsize=15)
plt.suptitle(f'Spatial Statistics for Selected Runs in "{store_name}"', fontsize=20)
return figFunction Producing Time Series Plots¶
Also show which dates have no available data values, as a rug plot.
def plot_timeseries(ds, data_var, store_name):
"""Plot the mean, min, max, and standard deviation values for several climate runs,
aggregated over lat/lon dimensions."""
num_members_to_plot = 4
member_names = ds.coords['member_id'].values[0:num_members_to_plot]
figWidth = 25
figHeight = 20
linewidth = 0.5
numPlotColumns = 1
fig, axs = plt.subplots(num_members_to_plot, numPlotColumns, figsize=(figWidth, figHeight))
for index in np.arange(num_members_to_plot):
mem_id = member_names[index]
data_slice = ds[data_var].sel(member_id=mem_id)
unit_string = ds[data_var].attrs['units']
min_vals = data_slice.min(dim = ['lat', 'lon'])
max_vals = data_slice.max(dim = ['lat', 'lon'])
mean_vals = data_slice.mean(dim = ['lat', 'lon'])
std_vals = data_slice.std(dim = ['lat', 'lon'])
missing_indexes = np.isnan(min_vals)
missing_times = ds.time[missing_indexes]
axs[index].plot(ds.time, max_vals, linewidth=linewidth, label='max', color='red')
axs[index].plot(ds.time, mean_vals, linewidth=linewidth, label='mean', color='black')
axs[index].fill_between(ds.time, (mean_vals - std_vals), (mean_vals + std_vals),
color='grey', linewidth=0, label='std', alpha=0.5)
axs[index].plot(ds.time, min_vals, linewidth=linewidth, label='min', color='blue')
ymin, ymax = axs[index].get_ylim()
rug_y = ymin + 0.01*(ymax-ymin)
axs[index].plot(missing_times, [rug_y]*len(missing_times), '|', color='m', label='missing')
axs[index].set_title(mem_id, fontsize=20)
axs[index].legend(loc='upper right')
axs[index].set_ylabel(unit_string)
plt.tight_layout(pad=10.2, w_pad=3.5, h_pad=3.5)
plt.suptitle(f'Temporal Statistics for Selected Runs in "{store_name}"', fontsize=20)
return figProduce Diagnostic Plots¶
Plot First, Middle, and Final Timesteps for Several Output Runs (less compute intensive)¶
%%time
# Plot using the Zarr Store obtained from an earlier step in the notebook.
figure = plot_first_mid_last(ds, data_var, store_name)
plt.show()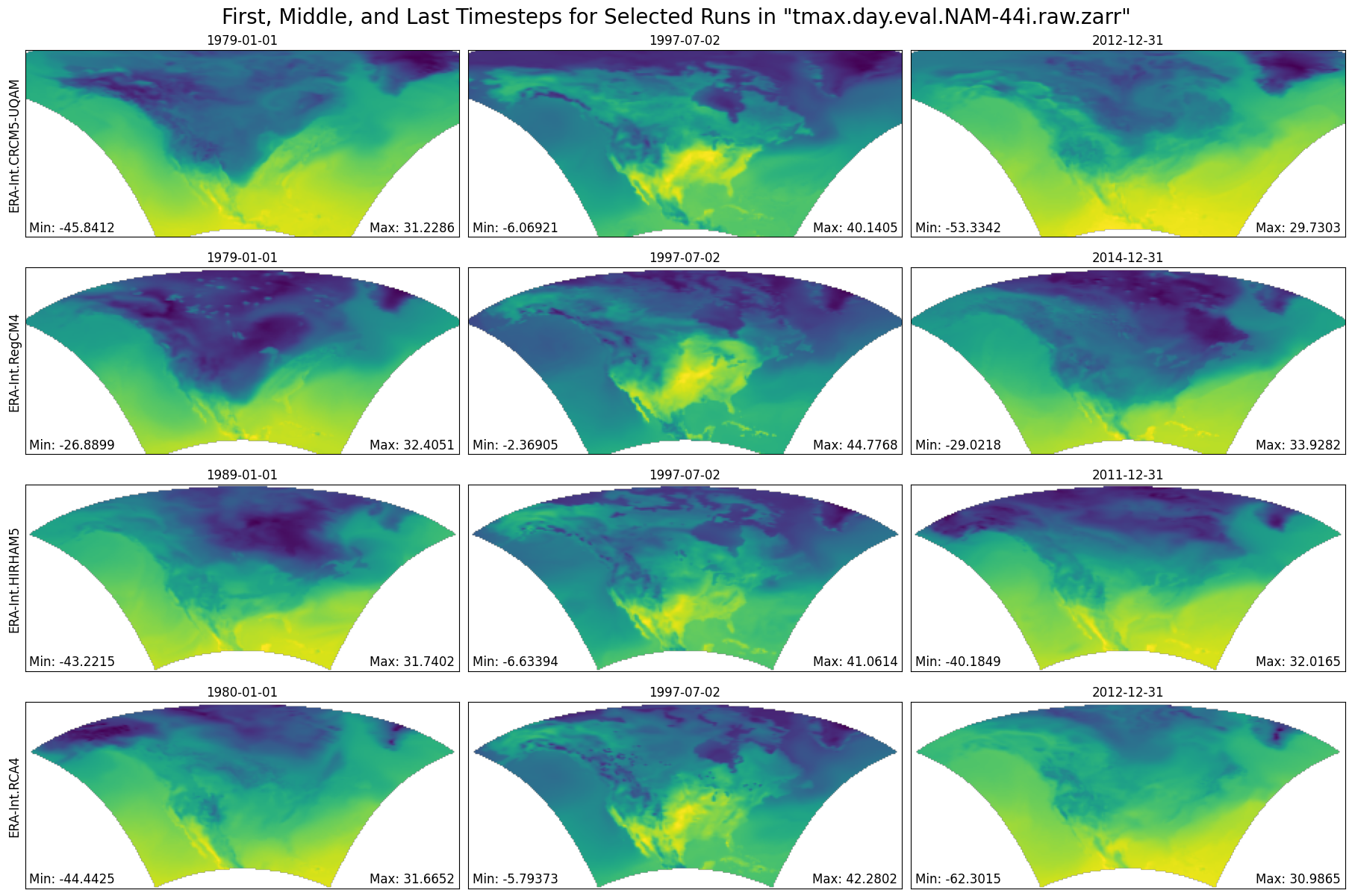
CPU times: user 2.48 s, sys: 221 ms, total: 2.7 s
Wall time: 44.7 s
cluster.close()